PipHelper retrieves sequence as well as annotation data from the UCSC Genome Browser and generates sequence, repeats, and exons files suitable for further processing by PipMaker and MultiPipMaker.
Repeats in the returned sequence have been "soft masked" using RepeatMasker as described in the UCSC Data and Downloads FAQ.
The application itself can be found here:
To use PipHelper, you need to choose an assembly, enter a position, and optionally choose an annotation.
The assemblies are pre-defined by the PipHelper server, and mirror the choices available at the UCSC Genome Browser. Assemblies are organized by clade and genome. For example, UCSC's hg17 can be found by choosing Vertebrate, Human, and then May 2004 (hg17). Similarly, apiMel2 can be found by choosing Insect, A. mellifera, and then Jan 2005 (apiMel2).
Position is entered as a range on a chromosome or scaffold. The format of the position is: chromosome:begin-end. A few examples of positions: chr11:1900000-3000000, SCAFFOLD100016:1-30000, GroupUn:47426601-47431800.
Annotation data can be optionally added by choosing an annotation from the dropdown menu. For example, one could choose to generate an exons file for a given position on the human genome from the RefSeq Genes.
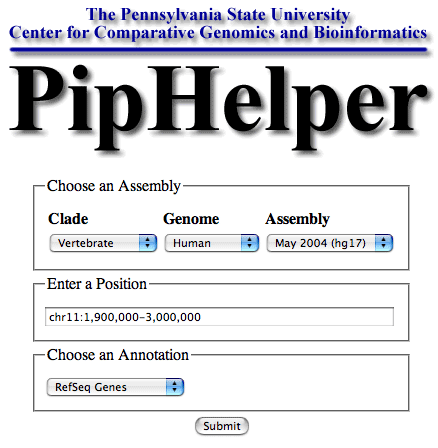
A screenshot of the PipHelper selection screen:

PipHelper generates text files which can be downloaded to your computer, and submitted to PipMaker and MultiPipMaker.
Sample output for hg17, chr11:100000-200000, RefSeq Genes
A screenshot of the PipHelper output screen: